Reinforcement Learning for Antibiotic Stewardship: Optimizing Prescribing Policies Under Antimicrobial Resistance Dynamics
1 Abstract
Antimicrobial resistance (AMR) threatens antibiotic effectiveness, but quantitatively evaluating the impact of stewardship strategies remains difficult using real-world data. We developed abx_amr_simulator, a Python simulation package that frames the problem of optimizing antibiotic prescribing as a Markov decision process. We used abx_amr_simulator to benchmark prescribing policies discovered by reinforcement learning agents against a clinically-realistic fixed prescribing rule across four sets of experiments of increasing complexity, where each set of experiments varied in type and degree of observed information degradation.
We quantitatively evaluated the quality of prescribing policies by assessing their ability to maximize the clinical benefit rendered to patients while also maintaining the clinical efficacy of antibiotics over the course of the simulations. For all scenarios, hierarchical RL agents outperformed flat RL agents, with flat RL agents discovering effective policies in only the simplest single-antibiotic setting, while hierarchical agents demonstrated effective learning across the range of scenarios tested. Adding memory to the RL agents did not reliably yield performance gains; in some scenarios, hierarchical recurrent agents outperformed hierarchical memoryless agents, but in other scenarios performance was comparable, suggesting that the marginal advantage of memory is context-dependent.
Patient heterogeneity and observable risk-stratification signals were major determinants of policy quality. When RL agents could differentiate higher- from lower-risk patients, they learned to preferentially treat high-risk patients and withhold treatment from low-risk patients — a triage strategy that substantially reduced prescribing and stabilized AMR at lower equilibria. Unexpectedly, policies learned under conditions of exaggerated risk stratification modestly outperformed accurate stratification. Under the most complex conditions — combining noisy patient observations, delayed AMR surveillance, and increased patient volume — hierarchical agents were able to converge to conservative low-AMR equilibria, while the fixed prescribing rule failed catastrophically by dring AMR levels to saturation.
Overall, our experiments demonstrate that hierarchical RL agents are capable of learning effective and farsighted antibiotic prescribing policies, and are robust against multiple types of information degradation.
2 Introduction
Rising antimicrobial resistance (AMR) is widely recognized as a major threat to global public health, adversely affecting the clinical care and outcomes of millions of patients worldwide by reducing the effectiveness of currently available antibiotics (WHO 2015). Recent estimates suggest that 4.95 million deaths globally in 2019 were associated with antibiotic-resistant bacterial infections, with a disproportionate burden borne by those living in resource-limited settings (Murray et al. 2022). In response, a range of interventions have been implemented to curb the emergence and spread of AMR (America (IDSA) 2011). In the United States, one of the primary strategies has been the establishment of antibiotic stewardship programs (ASPs) (Sutton and Ashley 2024), which are multidisciplinary efforts encompassing clinician education, dissemination of prescribing guidelines, compilation of antibiograms from microbiological data, and prospective review and optimization of antibiotic regimens, particularly in inpatient settings (Barlam et al. 2016).
Despite the widespread adoption of such programs, the quantitative evidence base evaluating their population-level impact remains limited. Many studies assessing the effectiveness of ASPs or related strategies have been constrained in geographic scope, duration, or both (Bertollo et al. 2018). A fundamental challenge to performing quantitative impact evaluation of these programs is the pervasive partial observability of key system components (Laxminarayan et al. 2013). Even in resource-rich settings where antibiotic prescription records are available, true selection pressure on pathogen populations is incompletely observed due to unmeasured sources of antibiotic exposure, including agricultural use and environmental contamination (Van Boeckel et al. 2015). Measurement of AMR itself presents an even greater challenge: while initiatives such as the WHO’s Global Antimicrobial Resistance and Use Surveillance System (GLASS) have sought to standardize surveillance, participation remains voluntary and coverage incomplete, and no universally adopted scalar metric of resistance at the community level yet exists (Organization 2022; Leth and Schultsz 2023). One commonly used proxy, the antibiogram, is subject to substantial biases, as it usually reflects only cultured clinical isolates from patients seeking care and is typically updated infrequently, often on an annual basis (Truong et al. 2021). Consequently, antibiograms provide an incomplete and temporally lagged view of underlying resistance dynamics.
Given these challenges, it is difficult to directly quantify the long-term effects of antibiotic stewardship program interventions using observational or interventional studies alone (Bertollo et al. 2018; Schweitzer et al. 2019). When real-world systems lack sufficient data to support direct modeling, simulation can offer a complementary framework in which researchers are able to directly and explicitly specify the underlying ground truth of key mechanisms, enabling controlled investigation of trade-offs that are difficult or impossible to observe in real-world settings.
In that spirit, the purpose of this work is to examine how varying levels of information degradation impact the performance of different antibiotic-prescribing algorithms. We used abx_amr_simulator, a Python simulation package designed to model the relationship between prescribing decisions and antimicrobial resistance dynamics, and specifically to capture the trade-off between immediate clinical benefit and long-term resistance management. A detailed description of the simulator’s architecture and functionality is provided in a separate manuscript (Lee and Blumberg 2026). We used the simulator to create a series of simulated scenarios of varying complexity, where observed characteristics of the environment were affected by different types of information degradation.
We then compared the performance of two types of prescribing algorithms in these scenarios: a ‘baseline’ fixed prescribing rule that emulates how real-world prescribers make decisions about antibiotic treatment, and reinforcement learning (RL) algorithms. Because the consequences of prescribing decisions unfold over time — treatment decisions made today can drive resistance that undermines antibiotic efficacy months later — antibiotic prescribing is fundamentally a sequential decision-making problem. Reinforcement learning is a subfield of machine learning focused on designing algorithms to optimize sequences of decisions in order to maximize cumulative long-term outcomes, making it a useful approach for this problem (see Section 3.1 for further details).
We note here that this application of simulation and reinforcement learning to the area of antimicrobial resistance is relatively novel. Prior simulation-based work in this domain has largely operated at the pathogen level, modeling evolutionary dynamics of resistance under drug pressure (King et al. 2025; Weaver et al. 2024). While these models have yielded important insights into resistance evolution, they do not capture the patient-level clinical dynamics — including treatment decisions, outcomes, and feedback between prescribing behavior and population-level resistance — that are central to evaluating antibiotic stewardship interventions. Separately, there has been growing interest in applying machine learning to AMR-related problems, including resistance prediction and empirical antibiotic selection (Haredasht et al. 2025; Harandi et al. 2025). However, these approaches predominantly utilize supervised learning to focus on prediction rather than optimization, and typically operate on static snapshots of patient data rather than modeling the dynamic, longitudinal consequences of prescribing decisions at the population level. To our knowledge, the only prior application of reinforcement learning to an AMR-related problem is Weaver et al. (Weaver et al. 2024), which uses RL to identify optimal treatment cycling strategies at the pathogen level. Our work differs fundamentally in that we apply RL to optimize clinical prescribing policy within a patient-level simulator, with the explicit goal of evaluating trade-offs between individual clinical outcomes and population-level resistance.
In this study, we show that RL algorithms are capable of optimizing antibiotic prescribing policies even under conditions of partial observability, and that they outperform a clinically-realistic fixed prescribing rule by being more far-sighted and more robust to information degradation. By comparing the performance of the RL algorithms against the fixed prescribing rule baseline, we demonstrate that our simulator enables quantitative comparative policy analysis, allowing us to characterize the magnitude and nature of gains achievable by adaptive prescribing strategies relative to static prescribing rules.
In the following sections, we describe our experimental design, including how we simulate synthetic patient populations, define the reward function, and vary the quality and timeliness of information available to the prescribing agent.
3 Materials and Methods
3.1 Antibiotic Prescribing as a Sequential Decision Problem
Optimizing antibiotic prescribing is a sequential decision-making problem under uncertainty: prescribing decisions affect both immediate patient outcomes and longer-term resistance dynamics, and the true underlying state of the system is rarely fully observable. Formally, such problems can be modeled as a Markov Decision Process (MDP) — a mathematical framework for sequential decision-making in which a decision-maker interacts with an environment over a series of timesteps. At each timestep, the decision-maker observes the current state of the environment, selects an action, and receives a numerical reward reflecting the quality of that action; the environment then transitions to a new state. A decision rule that maps observations to actions is referred to as a policy. The goal is to find a policy that maximizes the cumulative reward earned over the course of an episode (a single traversal of the environment from start to finish).
In the antibiotic prescribing context, each MDP component maps naturally to a clinical concept:
- State: the observable attributes of current cohort of patients and the available AMR surveillance information for each antibiotic.
- Action: the prescribing decision — which antibiotic, if any, to administer to the current patient.
- Reward: a numerical score calculated as a weighted sum of patient outcomes (whether the patient was treated, whether treatment succeeded, and whether adverse outcomes occurred) and community-wide resistance levels; see Section 3.2.4 for details.
- Transitions: how resistance levels and the patient population evolve following each prescribing decision.
A key feature of this problem is partial observability: the true infection status and resistance profile of each patient are unknown at the time of treatment. Rather than observing ground truth, the decision-maker sees only probabilistic estimates of these quantities — estimates that may be noisy, biased, or temporally lagged. Partial observability of this kind is a central feature of real-world antibiotic stewardship, and its effects are systematically varied across the experiment sets in this study.
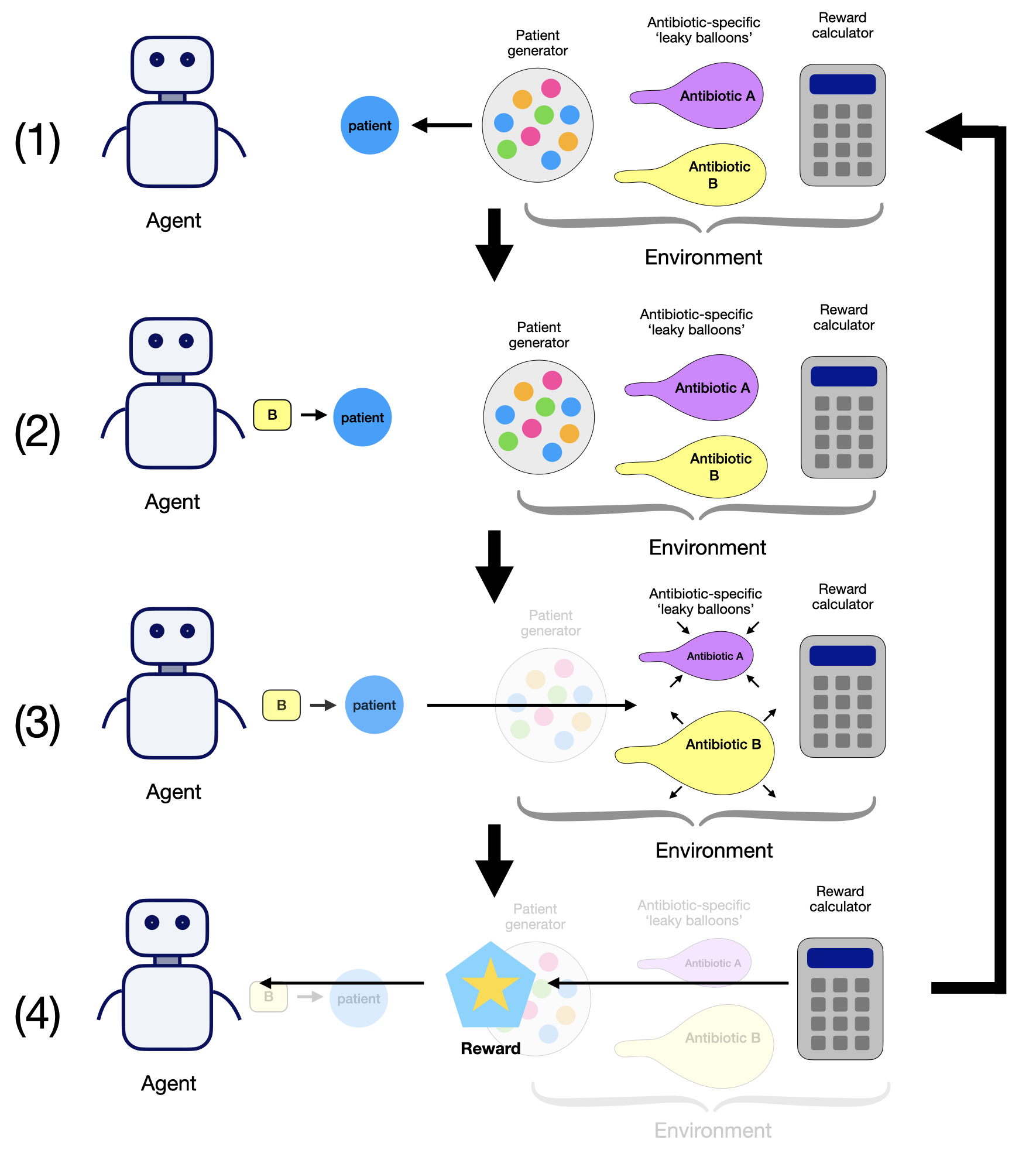
These experiments were conducted using abx_amr_simulator, a Python-based simulation framework that instantiates each component of the MDP described above: a PatientGenerator that presents patients to the prescribing agent at each timestep; an array of AMR_LeakyBalloon models that track and update resistance levels for each antibiotic; and a RewardCalculator that returns a numerical reward based on patient outcomes. A detailed description of the simulator’s architecture, functionality, and configuration options is available in the abx_amr_simulator GitHub documentation (Lee 2026) and companion preprint (Lee and Blumberg 2026); additional details on the simulator subcomponents are provided in Section S1 of Supplementary Material.
Because the MDP framework is agnostic to the choice of policy, any algorithm capable of selecting among a finite set of discrete actions and conforming to the Gymnasium API can be evaluated within this environment (Brockman et al. 2016; Towers et al. 2024).
In this study, we evaluate two classes of policies:
- A fixed prescribing rule (see Section 3.5), which emulates how real-world prescribers make antibiotic treatment decisions using observable patient and resistance information
- Reinforcement learning (RL) agents (see Section 3.3), which learn policies by interacting with the environment over many training episodes, using feedback from the reward signal to discover policies that maximize cumulative long-term reward. Unlike the fixed rule, RL agents are not pre-programmed with a specific decision logic; instead, they discover effective prescribing strategies through trial-and-error exploration.
For our RL agents, we used PPO (proximal policy optimization) implementations from the Python package stable-baselines3. The specific agent architectures used — flat memoryless, flat recurrent, hierarchical memoryless, and hierarchical recurrent — are described in Section 3.3.
A schematic overview of the simulator components and the agent–environment interaction loop is shown in Figure 1.

Note: this figure is reproduced from the companion software manuscript (Lee and Blumberg 2026).
3.2 Environment subcomponents
3.2.1 Patient Population and Observability
At each timestep, the environment uses the PatientGenerator subcomponent to generate a cohort of synthetic patients, which are characterized by attributes that determine their clinical profile and treatment outcomes. These attributes include probability of being infected, clinical benefit and failure multipliers, and spontaneous recovery probability. Attribute values can be set to constant values (creating homogenous patient populations), or drawn from a probability distribution (creating heterogeneous populations). These attribute values set by the PatientGenerator when creating patients determine the magnitude of the clinical reward vs. penalty earned by the agent for effective vs. ineffective treatment of the patient; however, the resistance profile of a potential infection is controlled by the array of AMR_LeakyBalloon models in the environment, and the resistance profile determines the likelihood that a given antibiotic treatment will be effective.
Patient attribute observations available to the RL agent can be manipulated by introducing noise or bias, or by concealing certain attributes altogether, therefore enabling exploration of prescribing strategies under varying levels of partial observability.
3.2.2 Antibiotic Resistance Dynamics
Antimicrobial resistance dynamics are modeled using the AMR_LeakyBalloon class, which tracks resistance to each antibiotic as a function of cumulative prescribing pressure. The model operates as a bounded accumulator: prescribing a given antibiotic increases the internal latent ‘pressure’, while in the absence of prescribing, this internal pressure decays over time as selection pressure is relieved — analogous to a balloon that inflates as air is pumped in, but slowly deflates when left alone. The observable resistance level is derived from this latent pressure via mapping through a sigmoid function, and represents the probability that a new infection at the current time is resistant to the given antibiotic. The characteristics of each antibiotic are defined by its resistance dynamics and side-effect profile. Resistance dynamics are specified via the AMR_LeakyBalloon parameters (e.g., how quickly resistance rises with increased prescribing and how quickly it decays when use is reduced), while side effects are defined by the probability and penalty magnitude of adverse events. The simulator also supports cross-resistance, allowing users to specify how prescribing one antibiotic influences resistance to others.
As with patient attributes, the fidelity of resistance level observations can be manipulated by introducing noise, bias, or delay, facilitating systematic investigation of prescribing strategies under imperfect resistance surveillance.
3.2.3 Treatment Decisions
At each timestep, the agent observes the current state of the environment, and based on what it observes, it makes an independent treatment decision for each patient: prescribe antibiotic A, prescribe antibiotic B, …, or withhold treatment. The information that the agent observes is probabilistic; that is, the agent cannot directly observe the true underlying attributes of the current cohort of patients (e.g., whether a patient is infected, and if so, what the resistance profile is of that infection). Instead, it sees noisy observations that encode uncertainty about these attributes, which we represent as probabilities over possible patient states.
The agent cannot prescribe multiple antibiotics to the same patient simultaneously, reflecting standard clinical practice of monotherapy (single-drug treatment) for straightforward infections. This constraint simplifies the decision space and aligns with real-world prescribing norms.
3.2.4 Reward Function
The reward function as calculated by the RewardCalculator balances immediate clinical outcomes against long-term antimicrobial resistance considerations. It consists of two components: the individual reward, which reflects the clinical benefit or failure of treatment decisions for the current cohort of patients, and the community reward, which penalizes high levels of antimicrobial resistance (AMR) across all antibiotics. These components are combined into a single scalar reward using a tunable weight parameter \(\lambda \in [0, 1]\), allowing the user to control the relative importance of individual versus community objectives. In all experiments reported in this manuscript, we fixed \(\lambda=0\), so agents were optimized using only individual clinical reward. This design allows us to test whether AMR-preserving behavior can emerge from long-horizon environment dynamics and policy architecture without explicit AMR reward shaping.
Importantly, although the agent acts under partial and noisy observations (Section Section 3.2.3), the simulator itself maintains access to the underlying ground-truth patient states. These latent states are used internally to generate clinical outcomes (e.g., whether a treatment succeeds or fails given the true infection and resistance status), and the reward is computed from these realized outcomes. The agent does not observe these latent variables or the reward-generating process directly; it only receives the scalar reward signal after each decision. This separation reflects real-world clinical settings, where treatment decisions are made under uncertainty, but outcomes (e.g., recovery or deterioration) are eventually observed and can be used to evaluate decisions.
The individual reward is calculated based on patient-specific attributes, antibiotic-specific properties, and discrete clinical outcomes (e.g., treatment success, failure, or adverse effects). The community reward, on the other hand, is based on the observed AMR levels of all antibiotics, ensuring consistency with realistic surveillance data rather than ground truth resistance levels. This design encourages agents to learn prescribing strategies that work within realistic informational constraints.
For mathematical definitions of the reward function, including equations, parameter descriptions, and details on stochastic modeling of clinical outcomes, see Section S2 of Supplementary Material.
3.3 Reinforcement learning agents
To evaluate how policy architecture affects antibiotic prescribing under uncertainty, we implemented four PPO-based agent classes using the stable-baselines3 Python library: flat memoryless PPO, flat recurrent PPO, hierarchical memoryless PPO, and hierarchical recurrent PPO.
The memoryless agents make each decision from the current observation only. By contrast, recurrent agents carry a short internal memory across timesteps, so recent observation history can influence the current decision. This distinction is important when AMR information is delayed or intermittently updated: agents that retain recent history may better approximate the latent current resistance state.
We also compared flat and hierarchical agent architectures. Flat agents choose a treatment action directly at each timestep. Hierarchical agents instead choose among higher-level options (also called workers; see (Sutton et al. 1999)), where each option encodes a clinically interpretable prescribing strategy over one or more steps. In our experiments, we used a ‘heuristic-worker’ option type, which uses fixed risk-based rules to choose treatment for each patient.
In this work, we define ‘heuristic workers’ as fixed, rule-based policies that map observable patient attributes to treatment decisions. Each worker computes an expected reward for available actions based on these observations and applies a predefined decision rule (e.g., “treat high-risk patients with antibiotic A,” “withhold treatment from low-risk patients”).
Importantly, these workers are not learned. Instead, they are hand-specified heuristics defined by parameterized rules. We constructed a library of such workers by instantiating these rules with different parameter values, yielding a diverse set of treatment strategies. In the single-antibiotic setting, the manager selects from a library of 9 heuristic workers, while in the two-antibiotic setting, it selects from 15 workers. Creating a range of options for the manager agent to choose from allows it more flexibility in optimizing its antibiotic prescribing strategy.
For our implementation of hierarchical PPO agents in this study, reinforcement learning operates at the level of the manager policy, which is trained using the PPO algorithm to learn which worker to deploy at each timestep. The manager learns to dynamically select which worker to deploy based on AMR state and population context, but the workers themselves remain fixed throughout training.
Our use of hierarchical agents was motivated by the long-horizon nature of AMR dynamics, in which current prescribing decisions can affect resistance and treatment utility many timesteps later; hierarchical reinforcement learning has been established as one among several RL algorithms that can effectively learn in environments affected by long-horizon credit assigment issues. (For more information, please see Section S3 of Supplementary Material, which discusses the option types and instances that were used in these experiments.)
3.4 Experimental Pipeline
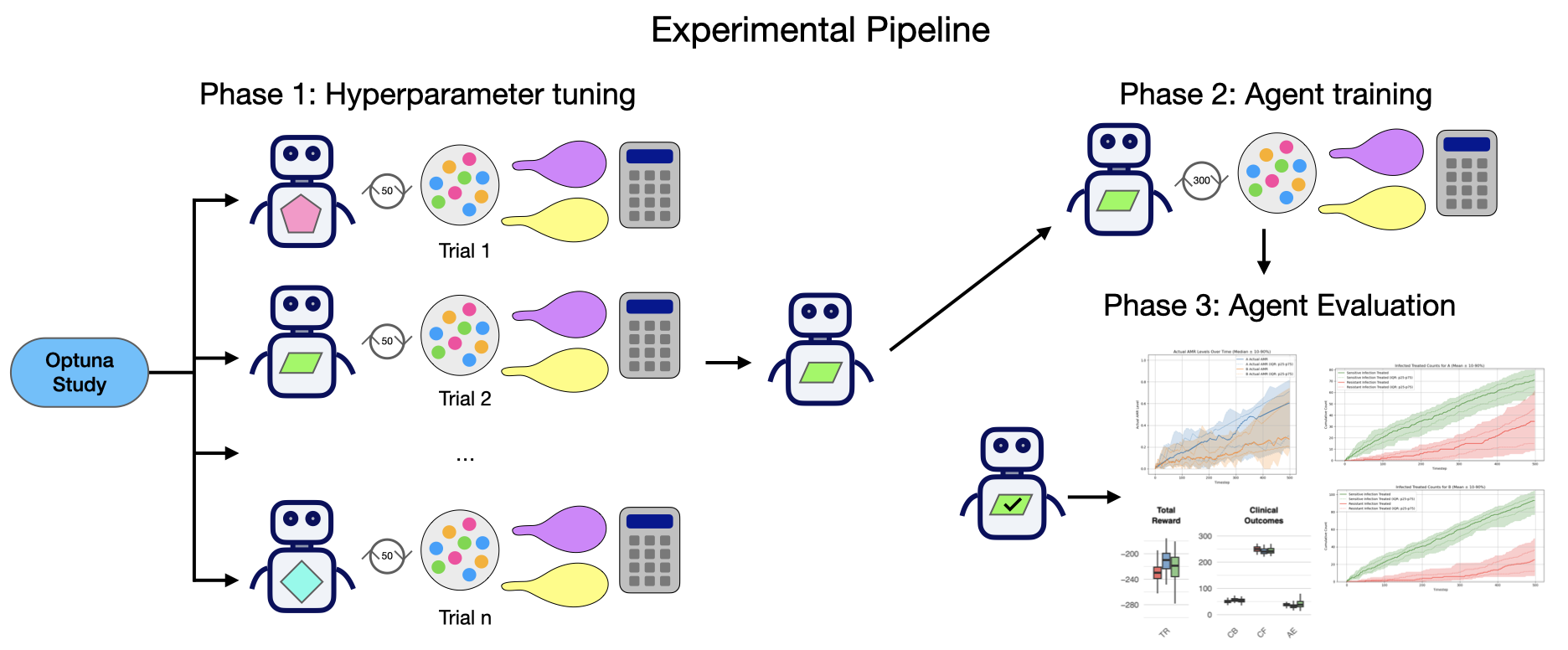
Effective reinforcement learning requires careful hyperparameter tuning. The abx_amr_simulator package uses the Optuna library (a hyperparameter optimization framework) for automated hyperparameter tuning (Akiba et al. 2019). This allows us to build modular end-to-end pipelines with three phases:
Tuning Phase: Optuna explores the hyperparameter search space, evaluating candidate configurations on preliminary environments. For each trial, the agent undergoes truncated training across 5 random seeds, with the study ultimately selecting the hyperparameter set that maximized a stability-adjusted objective (mean reward minus a modest variance penalty across seeds).
Training Phase: Multiple agent copies (20 per experiment) are initialized with optimized hyperparameters and trained on cloned target environments, where each environment is created from the same set of environmental parameters but initialized using a distinct random seed. This approach accounts for the empirical sensitivity of learned policies to initialization: we observed for certain scenarios and agent architectures, environmental stochasticity (particularly patient infection and resistance status across seeds) substantially influences policy acquisition, even with identical hyperparameters.
Evaluation Phase: Following training, each seed-specific trained agent executes 3 evaluative rollouts with logging enabled. During evaluation, we record three categories of cumulative metrics:
- Clinical Outcomes: patients experiencing benefit, failure, or adverse effects
- Patient Status: counts by true infection status (actually infected vs. not) and treatment received (any antibiotic given vs. none)
- Antibiotic Response: prescriptions per antibiotic, and sensitive vs. resistant infections treated
These metrics are agnostic to the decision algorithm, and can be computed for rollouts performed by RL agents, fixed prescribing rules, or any other type of strategy; these metrics therefore allow for fair, ‘apple-to-apples’ quantitative policy comparison. When the reward function is held constant while evaluating different policies, the raw overall reward achieved provides a valid shorthand for relative performance. However, examining cumulative count metrics remain essential for interpretation, since they show which outcome components (for example, clinical successes, failures, and adverse events) are driving differences in the overall reward earned by different strategies. All experiments in this study used the same reward function with fixed RewardCalculator hyperparameter values.
A schematic overview of the full experimental pipeline is shown in Figure 2.
3.5 Fixed Prescribing Rule Baseline
To evaluate whether learned policies provide value beyond static decision rules, we compared trained RL agents against a fixed prescribing baseline that performs no learning or adaptation.
Expected-reward (greedy): For each patient, compute the expected net clinical benefit (expected reward) of treating with each available antibiotic based on observed patient attributes and known scenario parameters. If no antibiotic has positive expected reward, withhold treatment. Otherwise, prescribe the antibiotic with the highest expected reward.
This fixed prescribing rule is intended to imitate how providers actually make decisions about antibiotic treatment in the real world, akin to static practice guidelines: they use available patient information and visible AMR data, but do not update behavior through experience. This makes them suitable non-learning comparators for isolating the incremental value of RL-based adaptation.
For fair comparison, fixed-rule policies are evaluated under the same rollout framework used for trained agents (multiple seeds used for evaluation), so performance differences reflect true differences in policy behavior rather than evaluation protocol artifacts.
3.6 Experiments: varying partial observability of patient attributes and AMR levels
Using the abx_amr_simulator package, we evaluated policies across three different antibiotic scenarios of increasing complexity: (1) a single-antibiotic scenario, (2) a two-antibiotic scenario without cross-resistance, and (3) a two-antibiotic scenario with moderate asymmetric cross-resistance. For the two-antibiotic scenarios, each antibiotic differs in
For each of these antibiotic scenarios, we varied patient-population structure and information quality across four sets of experiments, as summarized in Table 1. Populations were either homogeneous (fixed patient attribute values) or heterogeneous (attribute values sampled from Gaussian distributions), with some experiments introducing further heterogeneity with distinct high-risk and low-risk subpopulations. Importantly, the reward function was held constant across all experiment sets and antibiotic scenarios (with fixed RewardCalculator hyperparameter values), ensuring that observed performance differences reflect genuine differences in policy quality rather than reward design choices.
We now discuss the benchmark scenario construction, and describe the four experiment sets that progressively degrade observability and increase heterogeneity.
| Experiment Set | Patient Observability | AMR Observability | Patients / Timestep | Agent Architectures Evaluated | Hierarchical Agent Worker Type | Number of RL Experiments | Comparators | |
|---|---|---|---|---|---|---|---|---|
| 1 | Perfect Observability | Perfect | Perfect | 1 | Flat PPO; Hierarchical PPO | Heuristic Workers | 6 | Fixed prescribing rule: Expected reward - greedy |
| 2 | Delayed, Noisy, Biased AMR | Perfect | Delayed + noisy + biased | 1 | Flat PPO; Flat Recurrent PPO; Hierarchical PPO; Hierarchical Recurrent PPO | Heuristic Workers | 12 | Fixed prescribing rule: Expected reward - greedy |
| 3 | Heterogeneous Patient Attributes, Varied Bias | Biased observations | Perfect | 1 | Hierarchical PPO | Heuristic Workers | 9 | Fixed prescribing rule: Expected reward - greedy |
| 4 | Combined Observational Noise, Patient Heterogeneity | Noisy + biased + mixed visibility | Delayed + noisy | 10 | Hierarchical PPO; Hierarchical Recurrent PPO | Heuristic Workers | 6 | Fixed prescribing rule: Expected reward - greedy |
Benchmark scenario design and reference policies derived from fixed prescribing rules: In designing the three different antibiotic scenarios, we selected environment parameter values in order to produce nontrivial stewardship dynamics (i.e., situations that required policies with meaningful trade-offs, rather than trivial always-treat or never-treat behavior). In particular, for the homogeneous benchmark population we fixed baseline infection probability at \(p=0.7\), so treatment withholding is not trivially optimal and policies must balance near-term clinical benefit against longer-term resistance consequences.
In order to compute a sensible baseline to use as a comparator for the performance of the reinforcement learning agents, we rolled out the fixed prescribing rule ‘Expected Reward - Greedy’ as described in Section 3.5 in each benchmark scenario, under each of the conditions described in Table 1.
Experiment Set 1: Perfect Observability:
We first conducted a set of experiments in which the environment is fully observed: all patient attributes and antibiotic-specific AMR levels were presented to the agent without noise, bias, or delay, and all patient attributes are constant values and homogenous across all patients. These experiments serve as a best-case baseline, establishing the level of performance achievable by reinforcement learning agents and the baseline fixed prescribing rule when the true state of the system is fully known.
Experiment Set 2: Delayed, Noisy, Biased AMR:
In the second set of experiments, we introduced partial observability by injecting noise, bias, and temporal delay into the observed AMR levels, while maintaining perfect observability of patient attributes, where the patient population is again constant and homogeneous. This type of degraded information regarding AMR levels does resemble real-world clinical practice: as discussed in the Section 2, antibiograms provide an incomplete and typically delayed proxy for community resistance levels, requiring clinicians to make prescribing decisions using outdated and potentially biased information.
Introducing delayed updates to AMR observations renders the environment partially observable, as the prescribing policy now only has periodic access to updated resistance levels; otherwise, during inter-update periods, this information is stale. For this reason, in addition to memoryless PPO agents, we also evaluated recurrent PPO agents (both flat and hierarchical), which use LSTM (Long Short-Term Memory) networks to allow agents to integrate information over time. Intuitively, such agents may learn to maintain an internal estimate of latent resistance dynamics based on prior observations and prescribing outcomes, analogous to a clinician disregarding an outdated antibiogram, and instead forming informal beliefs regarding true current AMR levels based on accumulated experience.
Experiment Set 3: Heterogeneous Patient Attributes, Varied Bias:
In Experiment Set 3, we introduced heterogeneous patient populations with true differences in infection risk and treatment response while maintaining population-level averages matching the homogeneous baseline (Experiment Set 1). Patients were drawn from two subpopulations (high-risk and low-risk) with equal probability, where infection probability followed Gaussian distributions (\(\mu=0.84\) vs. \(\mu=0.56\), \(\sigma=0.1\)) rather than constant values. This creates ground-truth risk stratification that optimal policies should exploit.
We study the effect of information degradation by introducing bias into the observations provided to the agent, testing three conditions for each antibiotic scenario (see Table 2): (1) Accurate: true risk levels correctly observed, (2) Exaggerated: high-risk patients appear even higher risk, low-risk appear even lower, and (3) Compressed: risk differences appear smaller than they truly are. This isolates the cost of misperceiving patient heterogeneity when AMR information remains perfect and up-to-date.
| Perceived heterogeneity | Clinical interpretation | Agent Perception of Patient Risk |
|---|---|---|
| Accurate | Accurate risk stratification | True risk levels accurately observed |
| Exaggerated | Over-stratification | High-risk appear higher, low-risk appear lower |
| Compressed | Under-stratification | High-risk appear less high, low-risk appear less low |
Experiment Set 4: Combined Observational Noise, Patient Heterogeneity:
In the fourth and final set of experiments, we combine all sources of uncertainty from Experiment Sets 2 and 3: policies face noisy/biased patient observations alongside noisy/delayed AMR surveillance. We also substantially expand patient heterogeneity beyond Experiment Set 3. Whereas Experiment Set 3 introduced Gaussian variation only in infection probability (with other patient characteristics held constant within subpopulations), Experiment Set 4 samples all available patient attributes from Gaussian distributions. This creates both between-subpopulation differences (high-risk vs. low-risk mean parameter values) and within-subpopulation variation (\(\sigma=0.1\) for all attributes), more closely approximating real-world patient diversity.
Two additional design changes increase realism. First, we increase patient volume to 10 patients per timestep. Second, we implement differential observability: low-risk patients present with minimal workup (only 2 observable attributes: infection probability and spontaneous recovery probability), while high-risk patients undergo comprehensive assessment (all patient attributes are observable). This mirrors real-world clinical triage, where different patients have differential observability based on risk assessment.
Because delayed AMR observations render the environment partially observable over time, we evaluated both hierarchical PPO and hierarchical recurrent PPO agents to assess the impact of adding memory to the agent. This experiment set allowed us to evaluate how increasing noise and bias in both information streams, and delay in resistance information affected learned prescribing policies. We also then compared the performance of these learned policies to fixed prescribing rules that were operating under the same conditions of increased information degradation and increased patient volume.
4 Results
Full per-experiment AMR trajectories, cumulative clinical outcome plots, and summary statistics for all RL agents and fixed prescribing rules across all four experiment sets are provided in Supplementary Figures (RL Agents) and Supplementary Figures (Fixed Prescribing Rules), respectively.
4.1 Performance Metrics to Evaluate Policy Quality
Policy performance is evaluated using four classes of metrics:
- Total reward (TR) is the cumulative reward accrued over the simulation, reflecting the overall sum of treatment benefits, failures, and other rewards/penalties incurred by the sequence of decisions made by each policy rollout (see Supplementary Section S2).
- Clinical outcomes are summarized as counts of clinical benefit (CB), clinical failure (CF), and adverse events (AE).
- Patient status metrics track counts of patients by infection and treatment status: not infected and not treated (NI–NT), not infected but treated (NI–T), infected and not treated (I–NT), and infected and treated (I–T).
- Antibiotic response metrics summarize prescribing and effectiveness for each antibiotic, including total prescriptions (Rx), as well as counts of sensitive infections successfully treated (Sen–T) and resistant infections treated (Res–T).
All metrics are computed as cumulative values over full simulation rollouts and summarized across random seeds. Because the simulation environment is stochastic, repeated evaluations of the same policy yield variability in these outcomes. We therefore report distributions across seeds, visualized using boxplots that summarize the median and interquartile range.
4.2 Fixed Prescribing Rule: Expected Reward - Greedy Policy
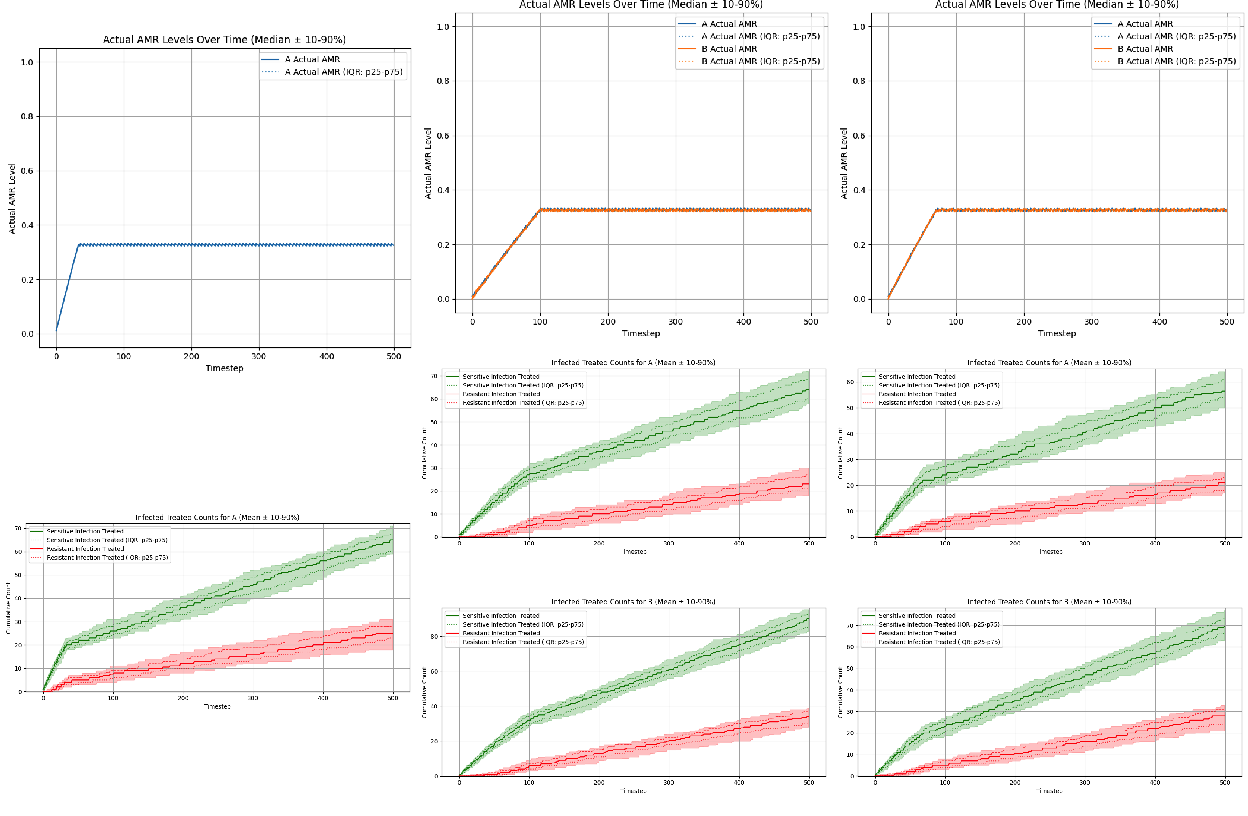
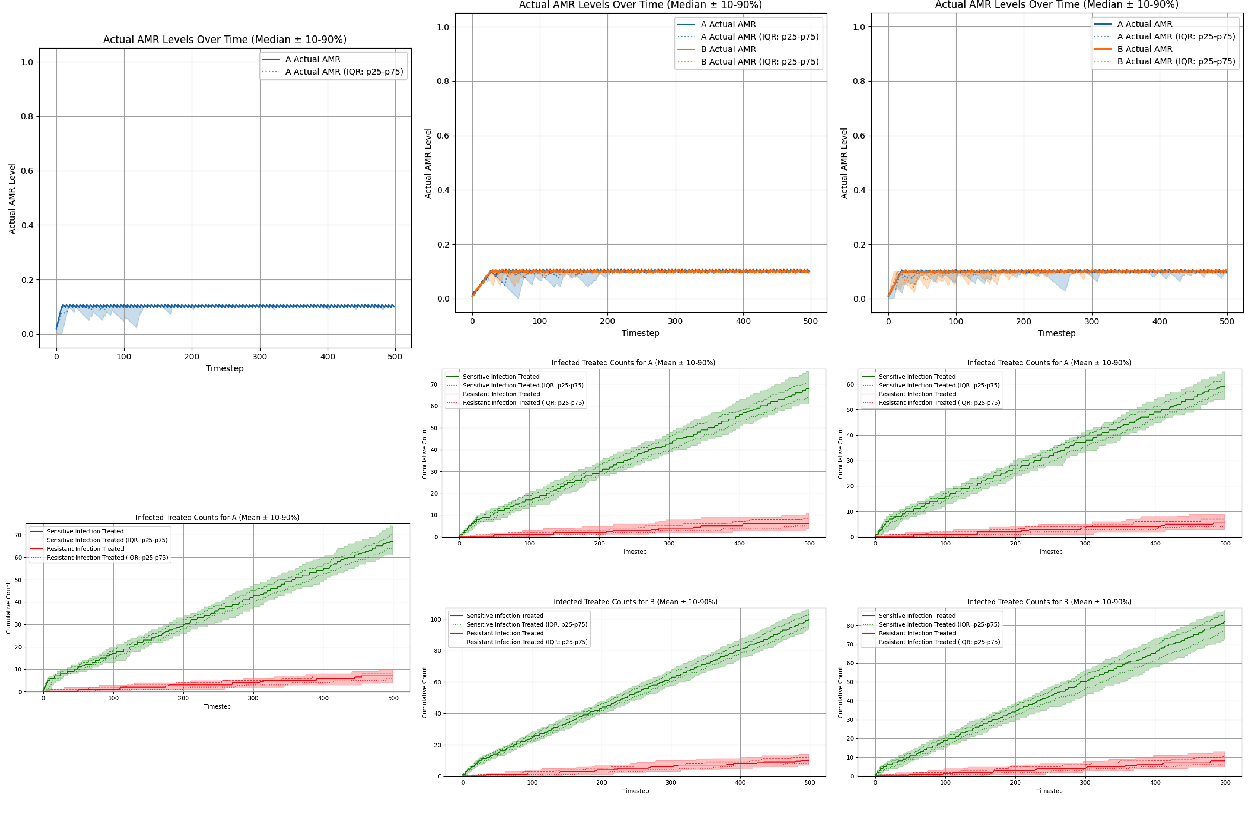
We used the fixed prescribing rule “Expected reward - greedy” as a baseline comparator for all three antibiotic scenarios under conditions of perfect observability. The fixed prescribing rule was rolled across 60 seeds per scenario. AMR trajectories (top panels) and cumulative clinical count metrics (bottom panels) are summarized in Figure 3. The cumulative clinical response plots allow us to evaluate the state of antibiotic efficacy throughout the episode by plotting counts of sensitive vs. resistant infections treated per antibiotic; sustained separation between these two curves, where sensitive counts remains above resistant counts, indicates preserved clinical efficacy over the episode. Conversely, if the curves intersect, or resistant counts rises above sensitive counts, this indicates a loss of clinical efficacy of the antibiotic.
- Single-antibiotic scenario: ‘Expected reward - greedy’ policy induces threshold-like prescribing that stabilizes AMR near an equilibrium of approximately 0.327 (Figure 3, left column).
- Two-antibiotic scenario (no cross-resistance): ‘Expected reward - greedy’ policy converges to essentially the same equilibria for each antibiotic: 0.327 for antibiotic A and 0.326 for antibiotic B (Figure 3, middle column).
- Two-antibiotic scenario (with cross-resistance): The ‘Expected reward - greedy’ policy again converges to essentially the same equilibria for each antibiotic: 0.326 for antibiotic A and 0.327 for antibiotic B (Figure 3, right column).
4.3 Experiment Set #1: Perfect Observability
Under perfect observability, we first asked a basic feasibility question: can reinforcement learning agents recover reasonable and effective prescribing policies when all patient attributes and AMR levels are fully and accurately observed at every timestep? This best-case setting therefore isolates the core learning challenge in these environments, which is tackling the long-horizon credit assignment problem.
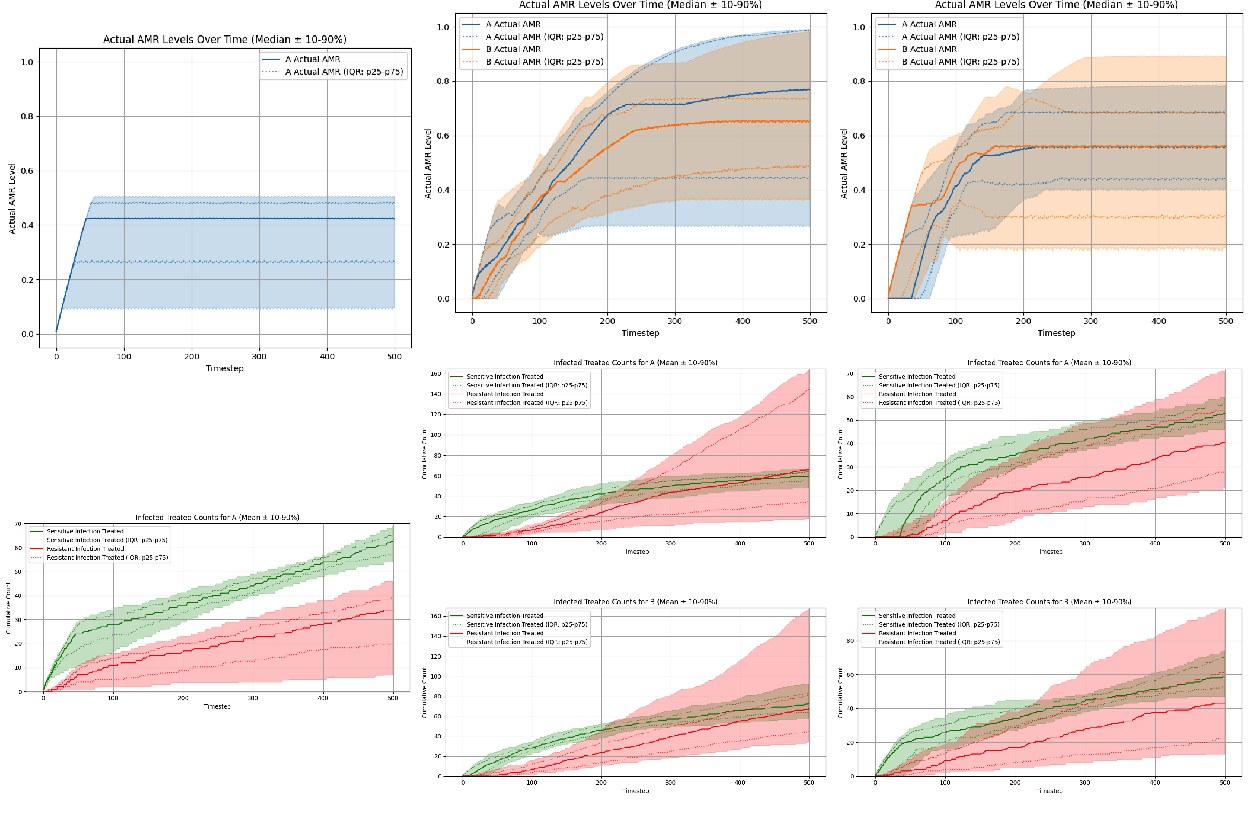
Flat PPO showed only limited success in this setting. In each of the three different scenarios, it learned policies that stabilized AMR levels at steady-state equilibria, but the learned equilibrium levels varied widely across seeds and did not consistently match the equilibria levels discovered by the ‘Expected Reward - Greedy’ fixed prescribing rule. In both two-antibiotic scenarios, the flat PPO performed particularly poorly: equilibrium AMR levels were highly variable across seeds, and cumulative sensitive-versus-resistant treated counts showed weak separation, indicating poorer preservation of antibiotic efficacy over the episode (Figure 4, middle and right columns).
As demonstrated in Figure 5, hierarchical PPO demonstrated much more robust learning across all three scenarios, with limited cross-seed variance. Interestingly, the policies discovered by hierarchical PPO specify significantly lower AMR equilibria levels compared to the ‘Expected Reward - Greedy’ fixed prescribing rule. We can see that the policies discovered by hierarchical PPO do outperform the fixed prescribing rule, with a more pronounced advantage for both two-antibiotic scenarios (Figure 6).
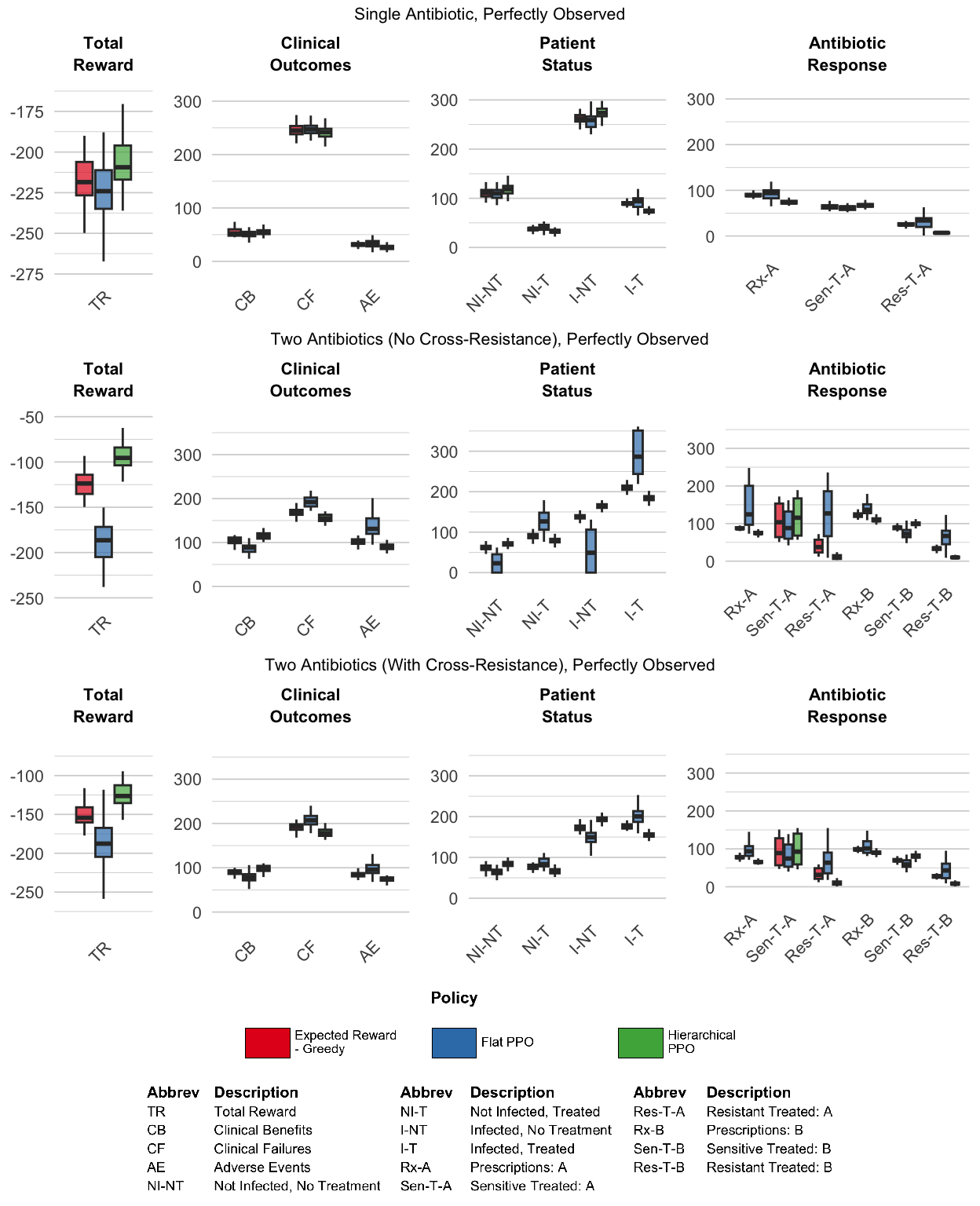
4.4 Experiment Set #2: Delayed, Noisy, Biased AMR
In Experiment Set 2, we inject noise, bias, and temporal delay into the observed AMR levels: AMR levels are updated every 90 timesteps, and the visible AMR signal is both noisy and biased (noise = 0.2, bias = -0.2). Figure 7 compares four PPO-based agents (flat memoryless, flat recurrent, hierarchical memoryless, hierarchical recurrent) against the ‘Expected Reward - Greedy’ fixed prescribing rule comparator across all three antibiotic scenarios.
The flat policies learned under degraded AMR information were qualitatively different from those learned under perfect observability, exhibiting a striking ‘on/off’ prescribing behavior driven by the AMR update cadence: policies concentrated treatment shortly after each AMR update, then pulled back during stale-information periods. Counterintuitively, the memoryless flat PPO agent expressed this pattern more cleanly than its recurrent counterpart, and outperformed it in both the single-antibiotic and two-antibiotic no cross-resistance scenarios (with similar performance in the two-antibiotic cross-resistance scenario). In the single-antibiotic scenario, the memoryless flat agent prescribed for a brief window following each AMR update, then ceased once the information grew sufficiently stale; the recurrent flat agent reduced prescribing during stale-information periods but did not stop as completely. In the two-antibiotic scenario without cross-resistance, this same contrast appeared as clean on/off alternation between antibiotics A and B for the memoryless flat agent, versus a softer reduction in use of the disfavored antibiotic for the recurrent flat agent. When cross-resistance was introduced, the distinction between the two prescribing patterns became less pronounced. Detailed prescribing time series and AMR trajectories are provided in Supplementary Figures (RL Agents).
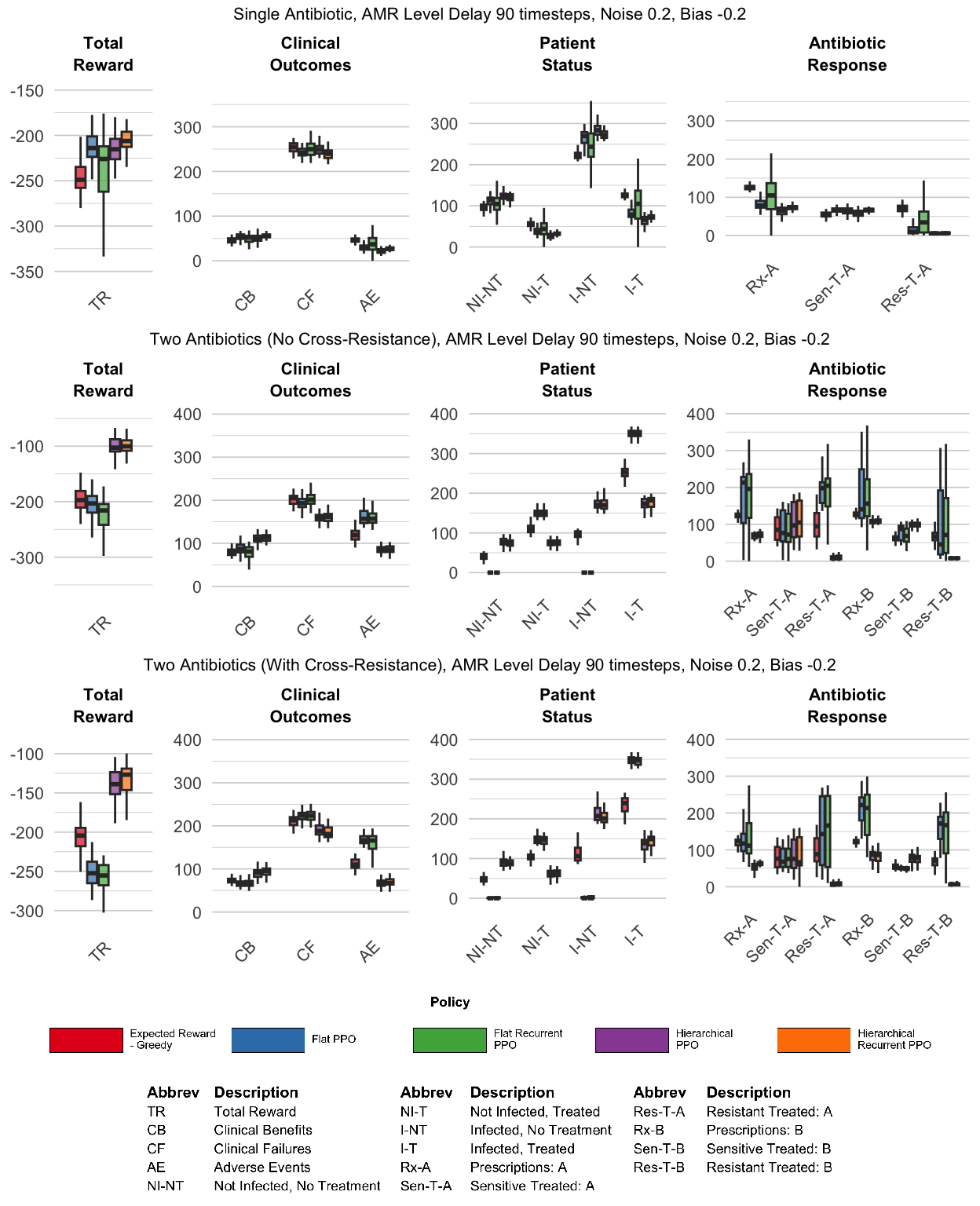
Unlike the flat agents, even under the conditions of degraded AMR information specified in Experiment Set 2, the hierarchical agents were able to learn policies that were qualitatively similar to the policy learned by the hierarchical PPO agent under conditions of perfect observability. Both HRL PPO and HRL recurrent PPO earned modestly lower overall reward than under perfect observability, but the performance gap was small (see Table 3). The hierarchical agents were able to recapitulate essentially the same AMR equilibria levels that were discovered under perfect-observability conditions, indicating that this architecture was able to robustly learn qualitatively similar policies even when AMR information was degraded. Furthermore, adding memory to the HRL architecture appeared to provide a small but consistent performance advantage. The AMR trajectories driven by the recurrent agent appeared steadier compared to those driven by the memoryless agent, consistent with our hypothesis that a persisted internal belief regarding AMR levels can improve decision-making when observations are stale.
| Agent (condition) | Single-abx | Two-abx (no CR) | Two-abx (CR) |
|---|---|---|---|
| HRL PPO (perfect observability, baseline) | -209.4 | -95.5 | -126.4 |
| HRL PPO (delayed/noisy/biased AMR) | -215.2 (-2.8%) | -102.7 (-7.6%) | -138.9 (-9.9%) |
| HRL recurrent PPO (delayed/noisy/biased AMR) | -206.2 (+1.5%) | -100.4 (-5.0%) | -126.8 (-0.3%) |
4.5 Experiment Set #3: Heterogeneous Patient Attributes, Varied Bias
In Experiment Set 3, we introduce true patient risk heterogeneity while keeping AMR observations perfect and up to date, then vary the fidelity with which policies perceive this heterogeneity across three conditions. We modeled two subpopulations (high-risk and low-risk) sampled with equal probability, with subpopulation parameters chosen so that population-average attribute values match the homogeneous baseline used in Experiment Set 1, allowing direct comparison to the results in that earlier experiment set.
Figure 8 shows results for the ‘Expected Reward - Greedy’ fixed prescribing rule (accurate risk stratification only) and HRL PPO (all three stratification conditions) across all three antibiotic scenarios. Introducing observable patient risk heterogeneity enabled HRL PPO agents to substantially improve their performance relative to the homogeneous baseline from Experiment Set 1, with gains observed across all scenarios and all stratification conditions. In contrast, the ‘Expected Reward - Greedy’ fixed prescribing rule showed mixed results, as shown in Table 4: performance degraded under accurate and compressed risk stratification and improved only modestly under exaggerated risk stratification, and these modest gains are substantially smaller in magnitude compared to the gains achieved by HRL PPO.
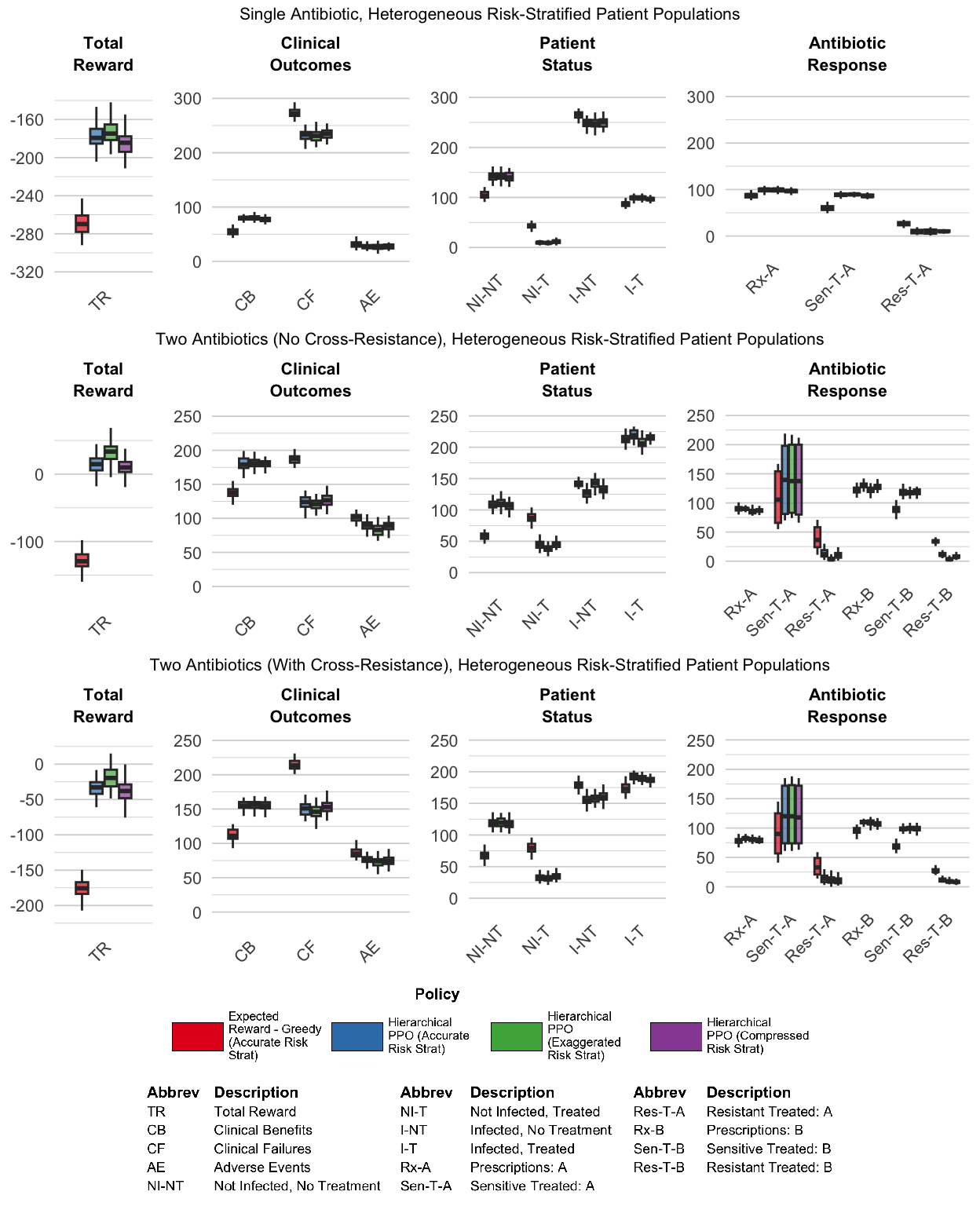
Table 4 also demonstrates that HRL PPO’s improvements over its Experiment Set 1 performance were substantial across all conditions, and the gains were largest in the two-antibiotic scenario without cross-resistance: HRL PPO improved by +114.5% under accurate risk stratification and by +135.0% under exaggerated risk stratification. The fixed prescribing rule underperformed or matched its own Experiment Set 1 performance in most conditions, achieving meaningful improvement only under exaggerated risk stratification in the two-antibiotic scenarios (+67.3% and +47.2% for ‘no cross-resistance’ and ‘with cross-resistance’ respectively).
| Condition | Single-abx | Two-abx (no CR) | Two-abx (CR) |
|---|---|---|---|
| Fixed prescribing rule | |||
| Baseline (Exp Set 1) | -218.5 | -123.8 | -154.4 |
| Accurate risk stratification | -269.8 (-23.5%) | -129.4 (-4.5%) | -175.7 (-13.8%) |
| Exaggerated risk stratification | -205.8 (+5.8%) | -40.5 (+67.3%) | -81.6 (+47.2%) |
| Compressed risk stratification | -270.8 (-24.0%) | -138.2 (-11.6%) | -185.7 (-20.2%) |
| HRL PPO | |||
| Baseline (Exp Set 1) | -209.4 | -95.5 | -126.3 |
| Accurate risk stratification | -179.3 (+14.4%) | +14.8 (+114.5%) | -32.9 (+74.0%) |
| Exaggerated risk stratification | -174.8 (+16.5%) | +33.4 (+135.0%) | -19.5 (+84.6%) |
| Compressed risk stratification | -184.4 (+11.9%) | -9.9 (+89.6%) | -38.0 (+70.0%) |
Interestingly, for both the HRL PPO agent and the fixed prescribing rule, performance under exaggerated risk stratification was better compared to performance under accurate risk stratification. Similarly, the HRL PPO agent and fixed prescribing rule also showed similar behavior for the compressed risk stratification condition: performance under compressed risk stratification was slightly worse than accurate stratification. The largest performance gains across all stratification conditions appeared in the two-antibiotic scenario without cross-resistance (Figure 8, middle panels). In the cross-resistance scenario, these gains were attenuated relative to the no cross-resistance case (Figure 8, right panels).
4.6 Experiment Set #4: Combined Observational Noise, Patient Heterogeneity
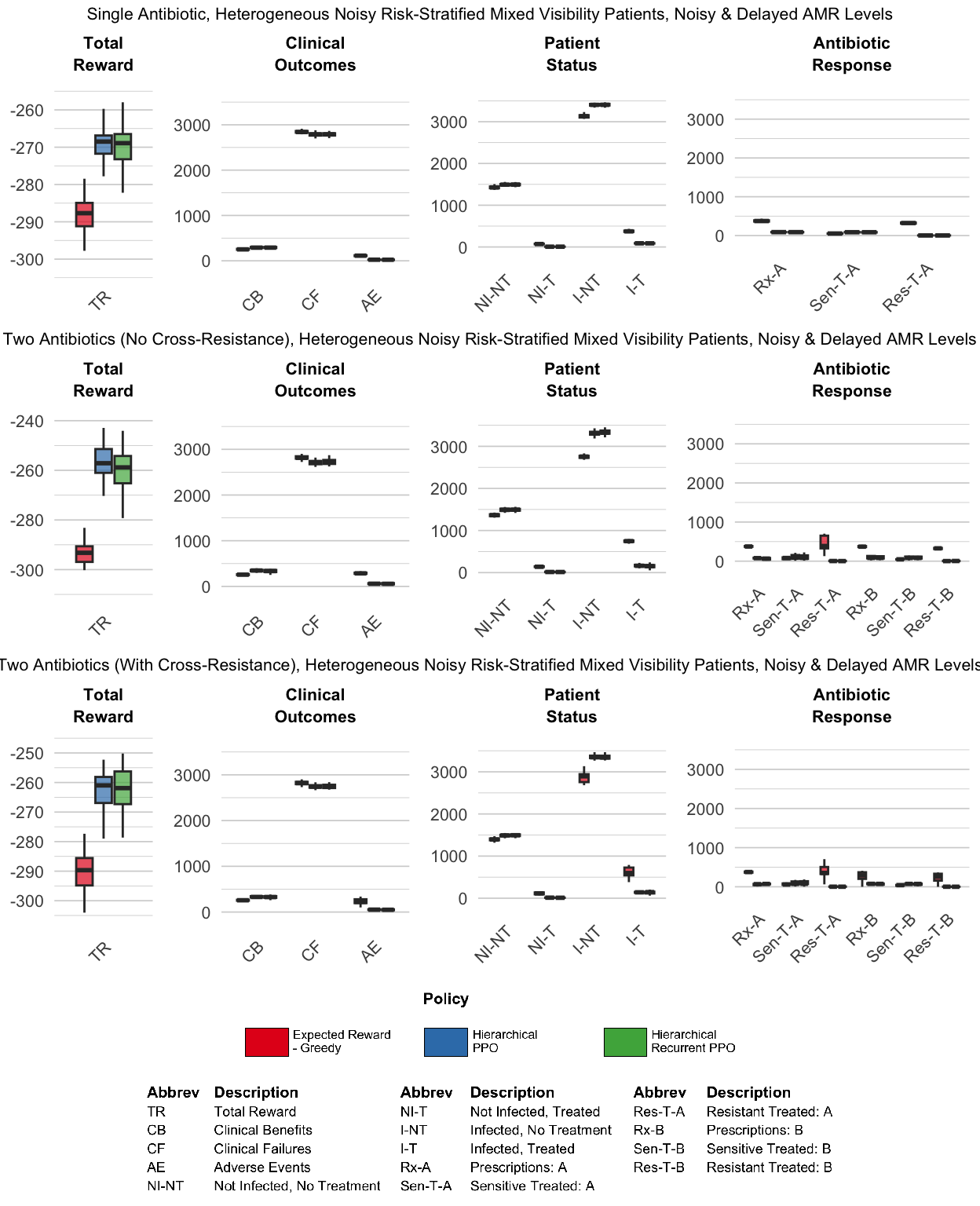
In Experiment Set 4, we combined both sources of uncertainty from Experiment Sets 2 and 3 and also increase patient volume: we specify noisy and biased patient observations, delayed and noisy AMR surveillance, and ten patients per timestep. We then compared the performance of hierarchical PPO and hierarchical recurrent PPO against the ‘Expected Reward - Greedy’ fixed prescribing rule for these scenarios with multiple sources of information degradation.
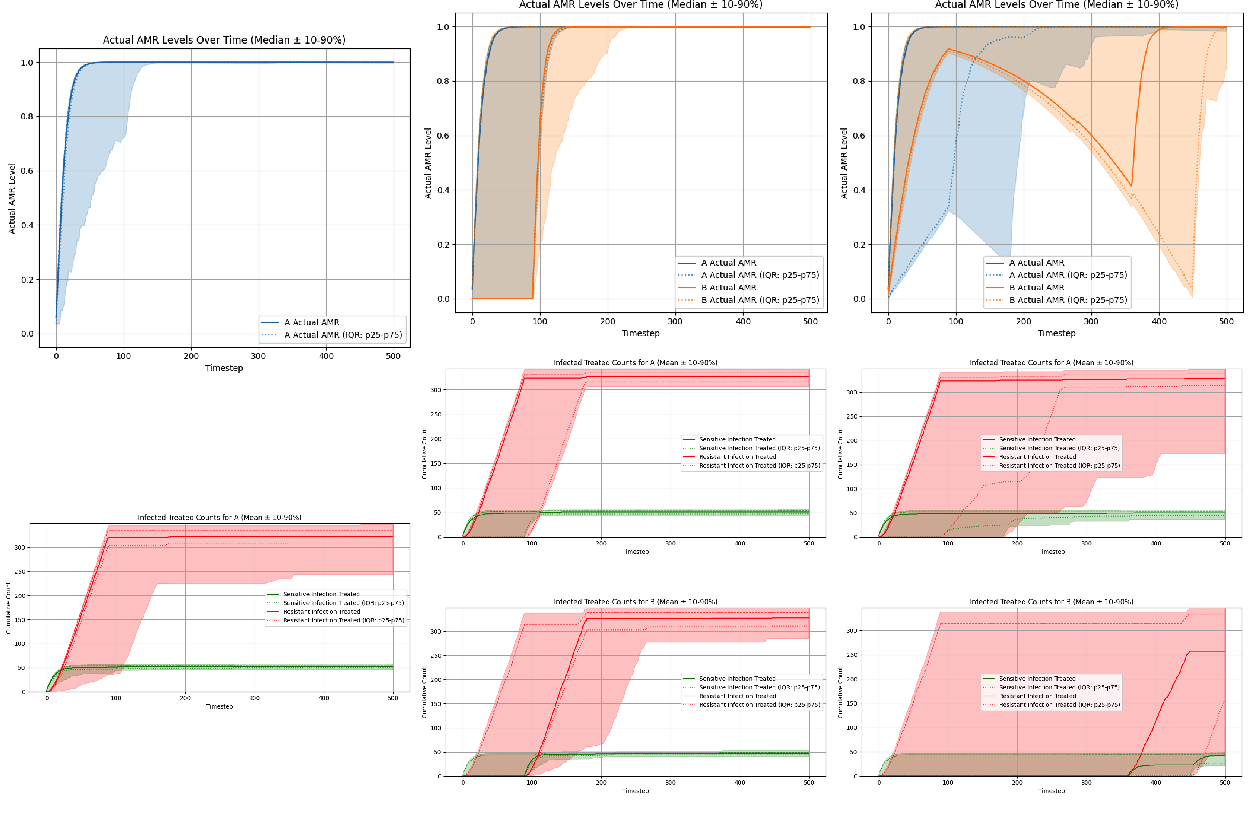
Both hierarchical architectures adopted markedly more conservative prescribing strategies than the “Expected Reward - Greedy” fixed prescribing rule comparator across all three antibiotic scenarios (Figure 10). Under the conditions of combined information degradation and increased patient volume, the “Expected Reward - Greedy” fixed prescribing rule showed it was no longer able to maintain steady-state equilibria for AMR levels, but instead exhibited monotonically increasing AMR levels over the episode, leading to depleted clinical efficacy of existing antibiotics, as demonstrated in Figure 9. These results demonstrate that in this set of experiments, the performance advantage of the hierarchical RL agents over the ‘Expected Reward - Greedy’ fixed prescribing rule is even more pronounced than in the prior sets of experiments. The hierarchical policies were not merely trading off short-term clinical outcomes for long-term stewardship: they achieved more clinical successes, fewer clinical failures, and significantly lower long-term AMR levels compared to the “Expected Reward - Greedy” fixed rule comparator.
Interestingly, for this set of experiments adding recurrence to the HRL PPO agent does not lead to improved performance, and in fact the hierarchical recurrent PPO agent demonstrates slightly worse performance compared to the hierarchical PPO agent. We revisit possible mechanisms for these results in Section 5.1. Figure 10 compares the cumulative performance metrics of the “Expected Reward - Greedy” fixed prescribing rule to the hierarchical PPO agents, both memoryless and recurrent.
5 Discussion
Across four experiment sets with varying kinds and degrees of information degradation, we found that reinforcement learning agents can discover effective antibiotic prescribing policies that dominate a clinically-realistic fixed prescribing rule, and that the learned policies are fairly robust against various types of information degradation. Agent architecture proved critical: flat PPO was sufficient only in the simplest single-antibiotic setting, while hierarchical temporal abstraction was required to handle the long-horizon credit assignment issues inherent in antibiotic prescribing in the context of antimicrobial resistance dynamics. Patient-level risk heterogeneity allowed hierarchical agents to substantially improve their learned policies over the fixed prescribing rule comparator, and under fully combined observational noise and increased patient volume, the fixed prescribing rule failed catastrophically while hierarchical agents converged to conservative low-AMR equilibria. We first examine each experiment set in turn, then draw cross-cutting conclusions about architecture and information structure, and close with study limitations and future directions.
5.1 Individual Experiment Set Findings
Experiment Set 1 (Perfect Observability): Under conditions of perfect observability and one patient per time step, the performance of the ‘Expected Reward - Greedy’ fixed prescribing rule and the HRL PPO agent were largely comparable, although the HRL PPO agent was able to modestly outperform the fixed prescribing rule in the two-antibiotic scenarios. In contrast, the flat PPO agent performed adequately only in the single-antibiotic scenario, but distinctly failed to learn effective policies in both two-antibiotic settings, even with complete state information. This suggests the failure mode is not informational but architectural: flat agents struggle with the long-horizon credit assignment required when prescribing decisions have delayed, coupled effects on future resistance levels. Switching to a hierarchical architecture — which decomposes the long-horizon problem into sequences of macro-actions — proved sufficient to recover effective policies across all three scenarios.
The comparable performance metrics between the fixed prescribing rule policy and the HRL PPO policy in this set of experiments suggest that when providers have perfect information, non-learning fixed rules can actually perform fairly well, and that the marginal advantage of adaptive learning offered by reinforcement learning agents is small.
Experiment Set 2 (Delayed, Noisy, Biased AMR): Under delayed, noisy, and biased AMR surveillance, the performance of the ‘Expected Reward - Greedy’ fixed prescribing rule and the flat PPO degrade significantly, although again the fixed prescribing rule actually outperforms the flat PPO in both of the two-antibiotic scenarios. However, the hierarchical agents demonstrated performance that was fairly robust to the degradation of the AMR information, exhibiting only a modest decrease in performance as measured by overall reward, and both agent architectures were able to learn policies that were qualitatively similar to the HRL PPO policy learned under perfect obervability. Adding memory led to a small but consistent performance boost across all three scenarios. This finding was consistent with our initial hypothesis: persisting an internal belief about latent AMR dynamics can improve decisions when observations are stale.
Experiment Set 3 (Heterogeneous Patient Attributes, Varied Bias): In Experiment Set 3, the marginal advantage of HRL agents over the fixed prescribing rule comparator is even more pronounced than in prior experiment sets. Introducing observable patient heterogeneity allows both types of algorithms to treat based on risk stratification, but HRL PPO exploits this far more effectively than the fixed prescribing rule. HRL PPO improved substantially on its Experiment Set 1 baseline across all scenarios and conditions; in the two-antibiotic (no cross-resistance) scenario, HRL PPO achieved positive overall reward under accurate and exaggerated risk stratification conditions, a notable contrast with the negative reward earned in every Experiment Set 1 scenario. Compressed risk stratification produced only modest degradation in performance by the HRL PPO agent, suggesting that the gains from heterogeneity are fairly robust to miscalibration as long as relative risk ordering is preserved — though the direction of miscalibration matters, with over-stratification preferable to under-stratification in the scenarios tested here.
The fixed prescribing rule, by contrast, performed worse than or comparable to its own Experiment Set 1 baseline under conditions of accurate and compressed risk stratification, and only showed modest improvement under conditions of exaggerated risk stratification. We suspect this improvement reflects a passive mechanism: under exaggerated stratification, low-risk patients appear even less likely to be infected than they truly are, reducing the expected reward for treating them and suppressing prescribing to that group — yielding ‘passive’ preservation of clinical efficacy. The larger and more universal gains achieved by HRL PPO suggest it learned a qualitatively different mechanism: a selective triage strategy that identified lower-risk patients across all three stratification conditions and withheld treatment from them to preserve long-term antibiotic efficacy.
For both HRL PPO and the fixed prescribing rule comparator, the largest gains across stratification conditions appeared in the two-antibiotic scenario with no cross-resistance. In this setting, the best strategy combines risk-stratified treatment with antibiotic rotation: the policies enacted by both the trained HRL PPO agent and the fixed prescribing rule tend to favor the antibiotic with lower current resistance, allowing the unused antibiotic’s resistance level to decay over time. This rotation does not appear as long contiguous blocks of single-antibiotic use (and is therefore not apparent in the plots in the Supplementary Figures), but instead emerges from rapid switching between antibiotics at the timestep level. As a result, both antibiotics are used throughout the episode, but in a coordinated manner that maintains lower resistance levels. In the cross-resistance scenario these gains are attenuated, since prescribing one antibiotic also drives up resistance to the other, reducing the benefit of such adaptive switching.
Experiment Set 4 (Combined Observational Noise, Patient Heterogeneity): In the most complex scenarios tested in this study, hierarchical PPO agents even more strongly dominate the fixed prescribing rule comparator, outperforming it across both clinical and stewardship metrics. Under conditions of combined information degradation and increased patient volume, the “Expected Reward - Greedy” fixed prescribing rule was no longer able to maintain lower steady-state equilibria for AMR levels, but instead ‘saturated’ them by driving the AMR levels up towards 100%, leading to depleted clinical efficacy of the existing antibiotics. We think a likely mechanism for this catastrophic failure of the fixed prescribing rule is due to the increased patient volume at each timestep; in the prior experiment sets, the fixed prescribing rule only had to treat one patient per timestep, meaning that its ability to drive up AMR levels too high before encountering the consequences was limited. In this set of experiments with 10 patients per timesteps, this natural bottleneck no longer exists, leading to the failure of the fixed prescribing rule to maintain long-term clinical efficacy of the antibiotics.
In contrast, both the HRL memoryless PPO and HRL recurrent PPO agents converged to conservative low-AMR equilibria that preserved efficacy of existing antibiotics throughout the episode. In contrast to the results from Experiment Set #2, adding memory did not appear to offer a performance advantage, and in fact the HRL memoryless PPO agent slightly outperformed the HRL recurrent PPO agent in the two-antibiotic scenarios. We suspect that the difference is that the scenarios tested in Experiment Set 2 were, in some ways, tailored to be the exact type of situation where memory would be most useful, i.e. AMR levels affected by noise and bias, with long delays between updates: therefore, an agent that was able to develop a good estimate of the true AMR levels and persist it between updates should be able to make better decisions compared to a memoryless agent. In contrast, the experiments here in Experiment Set #4 introduced additional sources of information degradation, and also increased the patient volume; for the scenarios tested in this set of experiments, it appears that the benefit of carrying an internal belief about AMR was offset by the cost of inferring it from increasingly noisy, heterogeneous observations.
5.2 Cross-Experiment Set Findings
Across all four experiment sets, the results confirm our central hypothesis: hierarchical RL agents learn antibiotic prescribing policies that match or dominate a clinically-realistic fixed prescribing rule, and these policies are robust to substantial information degradation. We note that while flat PPO was effective in the simplest setting, it failed to generalize across the multi-antibiotic scenarios; temporal abstraction via hierarchical RL was consistently required to handle the long-horizon credit assignment structure of antibiotic prescribing. We make no claim that hierarchical RL is the only viable approach for these environments, nor that it would scale without modification to substantially more complex settings (e.g., more antibiotics, larger patient cohorts, or richer resistance dynamics).
Three additional cross-cutting findings merit attention. First, and most surprisingly, risk over-stratification — where the agent perceives greater separation between high- and low-risk patients than actually exists — consistently yielded modestly better outcomes than accurate risk calibration; this was true for both the HRL PPO agent and the fixed prescribing rule comparator, although the advantage was more pronounced for the HRL PPO agent. One might expect that a miscalibrated perception of patient risk would degrade policy performance, but the opposite was true. We suspect two distinct mechanisms are at play: for the fixed prescribing rule, over-stratification further depresses the perceived expected reward for treating low-risk patients, producing the passive efficacy preservation described above; for HRL PPO, the effect appears more active — stronger perceived separation reinforces the agent’s learned tendency to withhold treatment from low-risk patients, enabling a more decisive selective triage strategy. This finding suggests that in settings where the cost of undertreating a low-risk patient is relatively low, a policy that errs toward over-stratification may be preferable to one calibrated to the true population risk distribution.
Second, observable patient risk heterogeneity enabled HRL PPO agents to learn a qualitatively different and highly selective prescribing strategy: agents learned to confidently withhold treatment from low-risk patients while concentrating prescribing on high-risk ones. The mechanism is that the manager converges to deploying moderate-threshold heuristic workers whose infection-probability thresholds naturally cleave the two subpopulations, reliably capturing high-risk patients while withholding treatment from almost all low-risk ones. This selective triage behavior appears to be the primary mechanism driving the large performance gains in Experiment Set 3 demonstrated by the HRL PPO agent architecture.
Third, adding memory appears to have mixed effects on performance; under conditions of delayed and noisy/biased AMR information, hierarchical recurrent agents had a small performance advantage compared to hierarchical memoryless agents. However, under the more complex scenarios tested in Experiment Set 4, adding memory did not provide an advantage, and the memoryless and recurrent hierarchical agents essentially achieved the same performance. This suggests that the added value of memory is context dependent.
Finally, the RL agents in these experiments achieved their stewardship outcomes despite being optimized using only individual clinical reward (\(\lambda=0\), with no explicit penalty for increasing community-level antimicrobial resistance). This indicates that when the environment itself creates long-term consequences for short-sighted prescribing — as AMR dynamics do — appropriately designed agent architectures can learn farsighted prescribing strategies without requiring explicit stewardship reward shaping.
5.3 Limitations
5.3.1 Abstraction of pathogen identity
Our simulation environment necessarily represents a simplified and idealized abstraction of real-world antibiotic prescribing and antimicrobial resistance dynamics. The most significant simplification involves our treatment of pathogens: we deliberately abstract away individual pathogen identity, even though in clinical practice pathogens serve as the natural unit for measuring antimicrobial resistance. Real-world antibiograms stratify resistance by pathogen species, tallying sensitive and resistant isolates separately for each organism; our simulator omits this pathogen-level granularity entirely.
We made this choice deliberately in order to distill clinical decision-making regarding antibiotic therapy to its most essential elements: (1) the probability that a given patient has a bacterial infection requiring antibiotic treatment, (2) the likelihood that a specific antibiotic will effectively treat that infection if present, and (3) the clinical trade-offs associated with prescribing. By reducing the decision to these core questions, we gain interpretability — both in how patients and AMR levels are represented, and in how the policies learned by RL agents can be understood. We also hypothesize that this simplification likely represents a theoretical upper bound on achievable clinical benefit; in a more stochastic, multi-pathogen environment, policy performance would likely degrade further.
5.3.2 Stationarity assumptions
In this study, we model both the patient population and the antimicrobial resistance response curve for each antibiotic as stationary entities — that is, patient attribute distributions and resistance dynamics remain constant throughout each episode and across episodes. In reality, neither is static: patient populations shift over time due to demographic change, seasonal variation in infection prevalence, and emergence of high-risk subpopulations, while resistance mechanisms can evolve in response to sustained selection pressure in ways that may be partially or fully irreversible. Policies learned under stationary assumptions may therefore be poorly calibrated to real-world settings where the underlying dynamics are drifting, and our results should be interpreted accordingly as characterizing performance under idealized stable conditions. Extensions to non-stationary dynamics are discussed in the Future Work section.
5.3.3 Single centralized prescriber
The simulator models a single, centralized prescriber with information about all patients and system-level AMR dynamics. This is unrealistic: real-world antibiotic prescribing involves multiple independent clinicians with limited visibility into their peers’ prescribing patterns and heterogeneous access to surveillance data. We plan to address this limitation in future work by extending the framework to support multi-agent, multi-locale environments where individual prescribers optimize locally with incomplete information (see Section 5.4).
5.3.4 Reinforcement learning algorithm selection
PPO was selected primarily for pragmatic reasons: it natively supports multidiscrete action spaces, as required for our multi-patient, multi-antibiotic setup, and is computationally tractable for the experimental scope of this study. Other RL algorithms or agent architectures may yield different performance characteristics. The modular design of the abx_amr_simulator package — which conforms to the Gymnasium API — allows interested users to implement and evaluate alternative agent classes beyond those tested here.
5.3.5 AMR resistance dynamic assumptions
The AMR_LeakyBalloon model makes specific assumptions about the shape of the antibiotic resistance response curve: latent resistance pressure is mapped to an observable AMR level through a sigmoid function, yielding a smooth, bounded accumulation and decay dynamic. While we believe this is a reasonable abstraction of resistance emergence and relief, the true functional form of resistance dynamics in real bacterial populations is not well characterized at the community level, and alternative response curve shapes could meaningfully alter the prescribing strategies learned by RL agents.
Additionally, our current implementation models cross-resistance as strictly positive — prescribing antibiotic A increases resistance to antibiotic B — whereas certain antibiotic combinations in fact exhibit collateral sensitivity, where use of one agent decreases resistance to another (Wang et al. 2025). Incorporating negative cross-resistance could meaningfully alter optimal prescribing strategies, potentially favoring cycling or mixing regimens that exploit these collateral effects.
Both of these limitations are mitigated by the modular architecture of the simulator. AMR dynamics are implemented through an abstract interface (AMRDynamicsBase), with AMR_LeakyBalloon as the concrete implementation used in this study. Users can provide alternative implementations of AMRDynamicsBase to specify different response curve shapes or cross-resistance structures, enabling systematic investigation of how resistance dynamic assumptions affect learned prescribing policies without requiring modification to the broader simulation framework.
5.3.6 Scope of findings and generalizability
Because the simulator is intentionally abstract, findings should not be interpreted as direct clinical policy recommendations. The framework is most useful for directional analysis under controlled conditions — specifically, quantifying how delayed, noisy, or biased antibiogram updates, miscalibrated patient-risk perception, and varying patient-level information quality affect prescribing decisions and the individual patient treatment benefit vs. antibiotic stewardship trade-off. Findings are hypothesis-generating and intended to inform the design of stewardship interventions and surveillance investments, not to prescribe specific clinical protocols.
5.4 Future Work
Several natural extensions of the abx_amr_simulator framework could substantially broaden its applicability and scientific scope.
5.4.1 Multi-agent, multi-locale environments
We plan to extend the framework to support multiple agents operating across geographically distinct locales. In this configuration, each locale would represent a separate community with its own AMR dynamics, and patients could migrate between locales, potentially spreading resistant infections. Individual agents would represent clinicians or healthcare facilities with limited visibility into peers’ prescribing practices — observing only their own cohort of patients and local antibiogram data. This extension would enable investigation of emergent dynamics in decentralized prescribing ecosystems, including how local optimization by individual agents affects community-level resistance, whether coordination mechanisms could improve collective outcomes, and how informational asymmetries across locales shape optimal strategies.
5.4.2 Non-stationary dynamics
Real-world patient populations and resistance mechanisms evolve over time, yet our current model treats both as stationary. Extending the simulator to support non-stationary dynamics would enable investigation of clinically important questions: How frequently should prescribing guidelines be updated to remain effective? What signals indicate that a guideline has become outdated? How much avoidable harm accumulates when policies fail to adapt to shifting patient demographics or pathogen ecology? We plan to address this by implementing time-varying patient attribute distributions to model demographic shifts, seasonal variation in infection prevalence, and emergence of high-risk subpopulations. We also plan to extend the leaky-balloon resistance dynamics to support regime-dependent behavior — for instance, modeling scenarios where sustained intensive antibiotic use creates irreversible changes in resistance mechanisms, such that the balloon accumulates a permanent baseline pressure that cannot be fully relieved even with extended periods of antibiotic restriction.
5.4.3 Upper bounds on personalized antibiogram prediction
Recent literature has explored machine learning approaches to predict patient-specific antibiotic susceptibilities from electronic medical record data, with the goal of improving empiric therapy selection (Al Mazrouei et al. 2025). While this body of work has grown substantially and spans a range of clinical settings, predictive models have generally achieved moderate discriminative performance (Sakagianni et al. 2023), leaving open a more fundamental question: even with perfect patient-level susceptibility prediction, how much collective clinical benefit could actually be achieved at the population level?
Our simulation framework is well positioned to address this question. By extending the patient model to include individual-level antibiotic susceptibility profiles, we can compare outcomes when RL agents have access to population-level antibiogram data alone versus perfect oracle knowledge of each patient’s true susceptibility — directly quantifying the theoretical ceiling on collective clinical benefit from personalized prediction under realistic conditions, including noisy observations, delayed resistance data, and heterogeneous patient populations. If this ceiling is high, it would justify continued investment in improving model performance despite currently modest discriminative metrics. If even perfect personalized prediction yields only marginal improvement over population-level antibiograms, it would suggest that resources might be more productively directed toward other stewardship interventions. We could further examine how this theoretical benefit varies across clinical contexts — including baseline resistance prevalence, population heterogeneity, and available antibiotic options — to identify settings where personalized prediction tools would provide the greatest value.
5.5 Conclusions
This study demonstrates how the abx_amr_simulator library can serve as a quantitative testbed for examining how information deficiencies shape antibiotic prescribing strategy performance under controlled conditions. Across four experiment sets of increasing complexity and worsening conditions of information degradation, we showed that hierarchical reinforcement learning agents can learn robust and effective prescribing policies that preserve antibiotic efficacy — outperforming a clinically-realistic fixed prescribing rule across clinical and stewardship metrics — without requiring explicit antimicrobial resistance penalties in the reward function. The framework supports directional analysis of factors including delayed or noisy antibiogram data, biased patient-risk perception, and differential access to patient-level clinical information, and their effects on the individual patient treatment benefit vs. antibiotic stewardship trade-off inherent in antibiotic prescribing. While findings are not intended as direct clinical guidance, they provide empirically grounded, hypothesis-generating insights that can inform stewardship program design, surveillance investment priorities, and the value of improving clinical risk assessment tools.
5.6 Acknowledgments
The authors thank Daniel De la Rosa Martinez and Paula Weidemueller for their helpful feedback on this work.
7 Funding
This work was supported by the CDC MInD-Healthcare (Modeling Infectious Diseases in Healthcare) grant U01CK000590.